Microbatch ingestion using Spark Streaming and Cassandra
In this post you’ll learn how to ingest data files in near real time using Spark Structured Streaming framework and storing them in Cassandra database. Spark Structured Streaming allows the programmer to implement the streaming solutions by means of dataframes instead of RDDs. This makes the development much more agile while preserving the performance thanks to the Catalyst optimiser, which runs as a compiler hidden to the programmer.
Spark Structured Streaming doesn’t make real time but near real time this means that the latencies are higher in comparison to real time. However, for many applications near-real time is more than enough and by using Spark the programmer has access to very powerful data transformations that can be done easily if we compare to use other frameworks such as Flink or Kafka Streams.
In Spark Structured Streaming the concept of sink determines how to finally put data into a destination. Because of not all databases or destinations have a sink, this tutorial shows an alternative way to ingest little batches of data into the final destination. A very common case is ingesting data to Cassandra as this database offers a scalable and redundant solution while being able to tune the degree of consistency. (Remember the CAP theorem).
Configuration of the environment
Let’s first configure the environment. Basically we need to download and configure Spark, Jupyter Notebooks and Cassandra. For Cassandra a straightforward way will be used as the tutorial is for testing cases.
Install Spark
You can find many ways or variants to install it. For this post the following three instructions are for MacOS.
brew cask install java brew install scala brew install apache-spark
Verify that you have Spark Scala and Pyspark installed.
spark-shell (quit from the interactive session and then run pyspark) pyspark
Add the spark connector package to operate with Cassandra database.
/usr/local/opt/apache-spark/libexec/conf/spark-defaults.conf
spark.jars.packages com.datastax.spark:spark-cassandra-connector_2.12:3.0.0-beta
Check the package is loaded correctly.
/usr/local/opt/apache-spark/libexec/bin/pyspark
You will see how the package is downloaded and loaded to spark correctly.
Install Jupyter Notebooks
pip3 install jupyterlab pip3 install notebook
jupyter notebook --port=80
Install Docker Desktop
To install Cassandra we’ll be using a ready to use distribution from Datastax. A requirement is to install first docker and kubernetes. But we’ll install those also very easily for local development.
Go to this website and download the Docker Desktop application: https://www.docker.com/products/docker-desktop
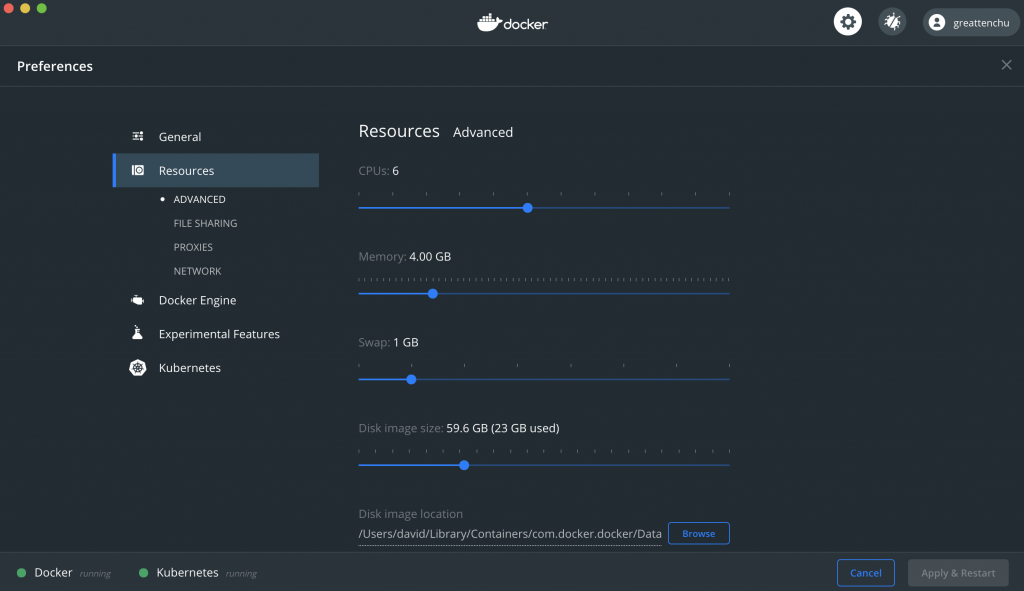
The visual aspect should be something like the image below.
Assign it 4 GB of RAM.
Install Cassandra
Datastax offers premium support for Cassandra. This is a very common case in which a private company adds a premium tier to an open source project. Go to their website and download Datastax Desktop.
https://downloads.datastax.com/#desktop
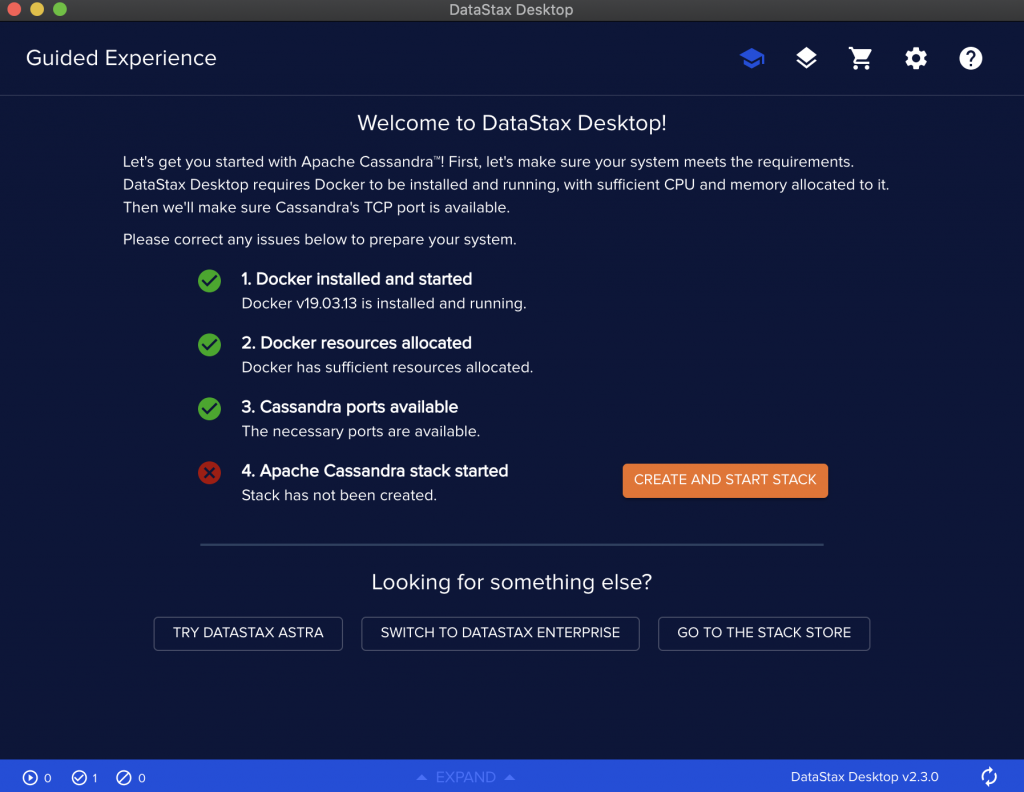
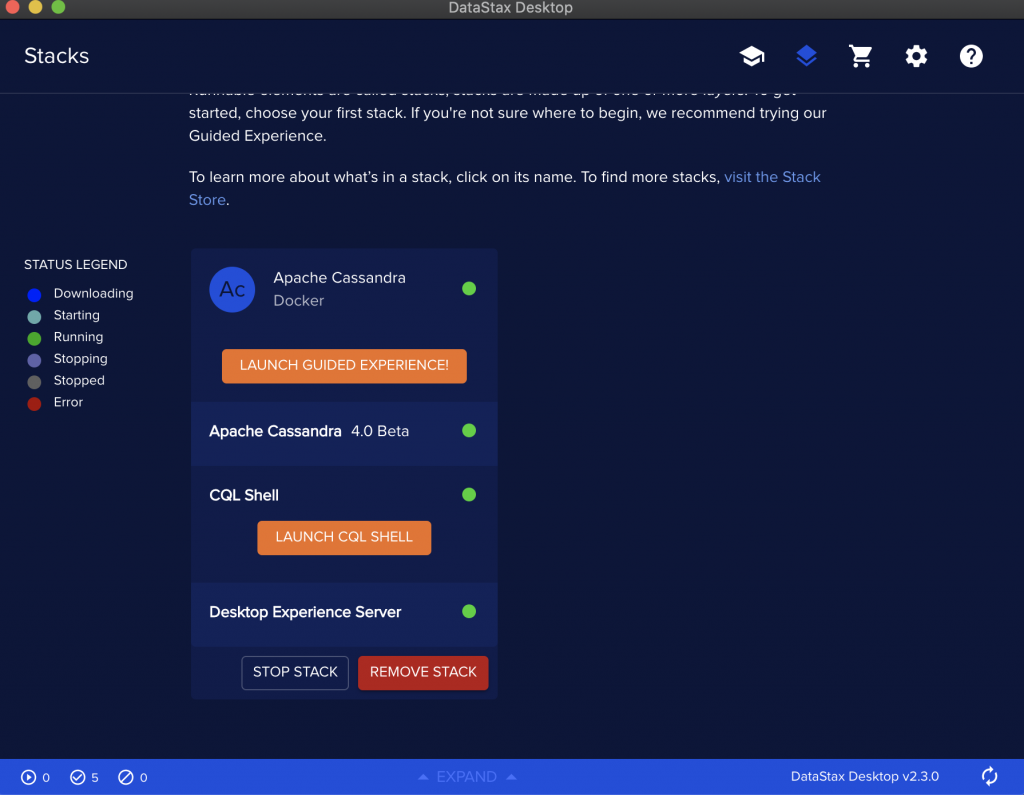
Make sure you meet the requirements and then create a stack to have an instance of Cassandra running on top of Kubernetes.
Use the Launch guided experience.
Preparing the database
Now it’¡s time to prepare the database. First create a keyspace and inside that keyspace create a table.
Create the keyspace and the table:
CREATE KEYSPACE consumptions WITH replication = {'class': 'SimpleStrategy', 'replication_factor' : 1};
CREATE TABLE c1( consumerID text PRIMARY KEY, date text, timevariation varint, measure1 varint, measure2 varint, filepath text );
Coding the application
The application looks very simple but it’s enough to deploy a system that ingests new files from a directory and loads their contents to Cassandra. Bear in mind that Spark is doing and keep running two processes in the background. One that detects and loads into dataframes new files, the second one is the one to load those micro batches to Cassandra.
Thanks to having the loaded data inside of a dataframe it’s very convenient to apply all transformations in between reading from the file and storing to Cassandra.
import findspark
findspark.init()
import pyspark
from pyspark.sql import SparkSession
from pyspark.sql.functions import expr
from pyspark.sql.functions import *
from pyspark.sql.types import StructType, StructField, IntegerType, FloatType, StringType, BooleanType,TimestampType, DateType
print("start")
##################################################################################################################
def write_to_cassandra(target_df, batch_id):
print("hi write cassandra")
df=target_df.selectExpr("cast(_c0 as string)consumerid", "cast(_c1 as string)date", "cast(_c2 as int)timevariation", "cast(_c3 as string)measure1", "cast(_c4 as string)measure2","filepath" )
df.write \
.format("org.apache.spark.sql.cassandra") \
.option("keyspace", "consumptions") \
.option("table", "c1") \
.mode("append") \
.save()
df.show()
print ("end write cassandra")
################################################################################################################
spark = SparkSession \
.builder \
.master("local[3]") \
.appName("Write to Cassandra") \
.config("spark.streaming.stopGracefullyOnShutdown", "true") \
.config("spark.sql.shuffle.partitions", 2) \
.config("spark.cassandra.connection.host", "localhost") \
.config("spark.cassandra.connection.port", "9042") \
.config("spark.sql.extensions", "com.datastax.spark.connector.CassandraSparkExtensions") \
.config("spark.sql.catalog.lh", "com.datastax.spark.connector.datasource.CassandraCatalog") \
.getOrCreate()
userSchema = StructType().add("_c0", "string").add("_c1", "string").add("_c2", "integer").add("_c3", "integer").add("_c4", "integer")
raw_df = spark.readStream \
.format("csv") \
.option("path", "/Users/david/bigdata/sparkstreamingfiles") \
.option("maxFilesPerTrigger", 1) \
.schema(userSchema) \
.load()
raw_df = raw_df.withColumn("filepath", input_file_name() )
output_query = raw_df.writeStream \
.foreachBatch(write_to_cassandra) \
.outputMode("update") \
.option("checkpointLocation", "chk-point-dir") \
.trigger(processingTime="1 minute") \
.start()
There are different ways to track changes and load them with Spark Streaming. For this example can investigate what’s the behaviour when restarting the SparkSession object (which runs in the master node). Although the SparkSession is deleted, Spark Streaming keeps a log of all inserted files with some metadata as an offset for the ongoing job. In this code, this is the chk-point-dir. Go to this directory in your machine and check that you have different folders that contain as many folders as files there have been loaded. Try to erase some of those subfolders and see how spark reload that information into Cassandra. However, as the operation acts and an upsert, if no modifications are set into the files the final data will remain the same in Cassandra.
Testing the system
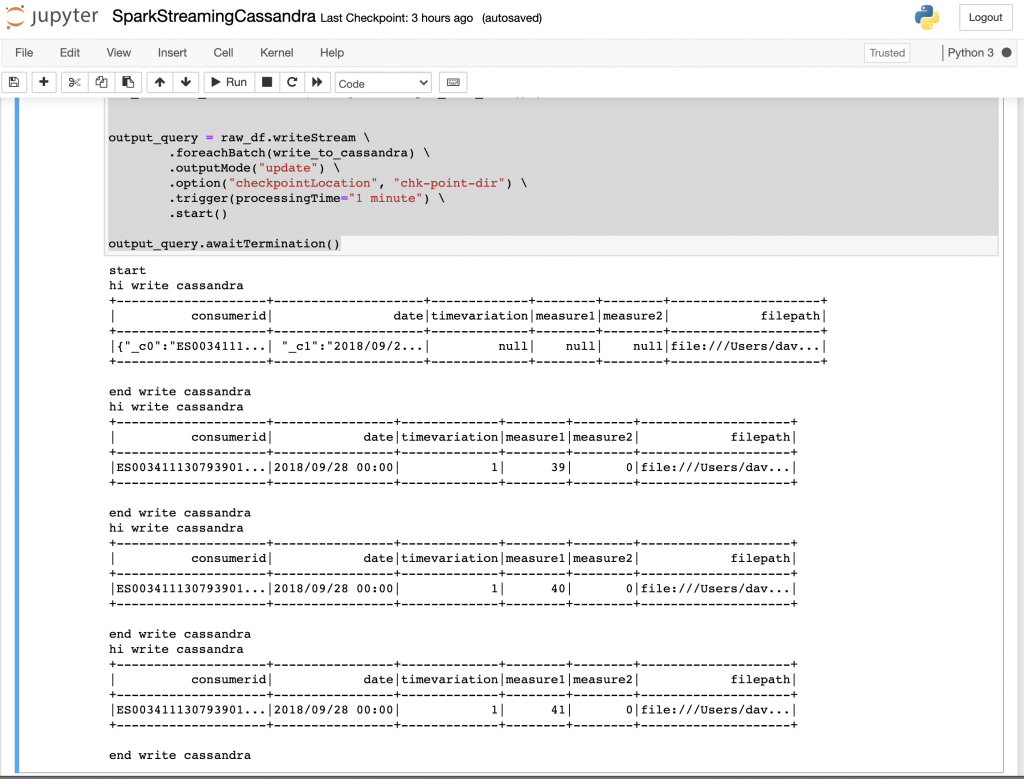
Press shift and intro to run the notebook block. The expected output should be something like:
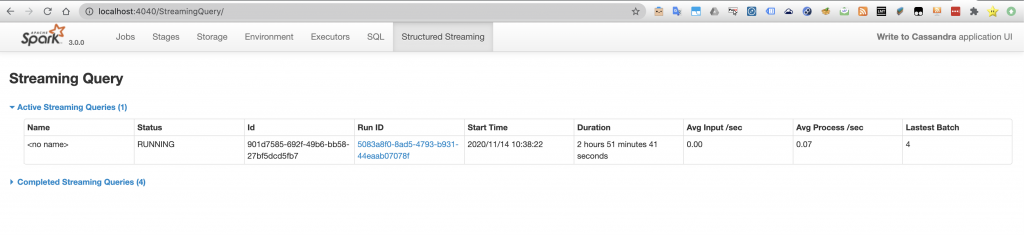
An interesting functionality is that because of the job was a streaming job, when going to the Spark UI there will be a tab for information related to structured streaming jobs.
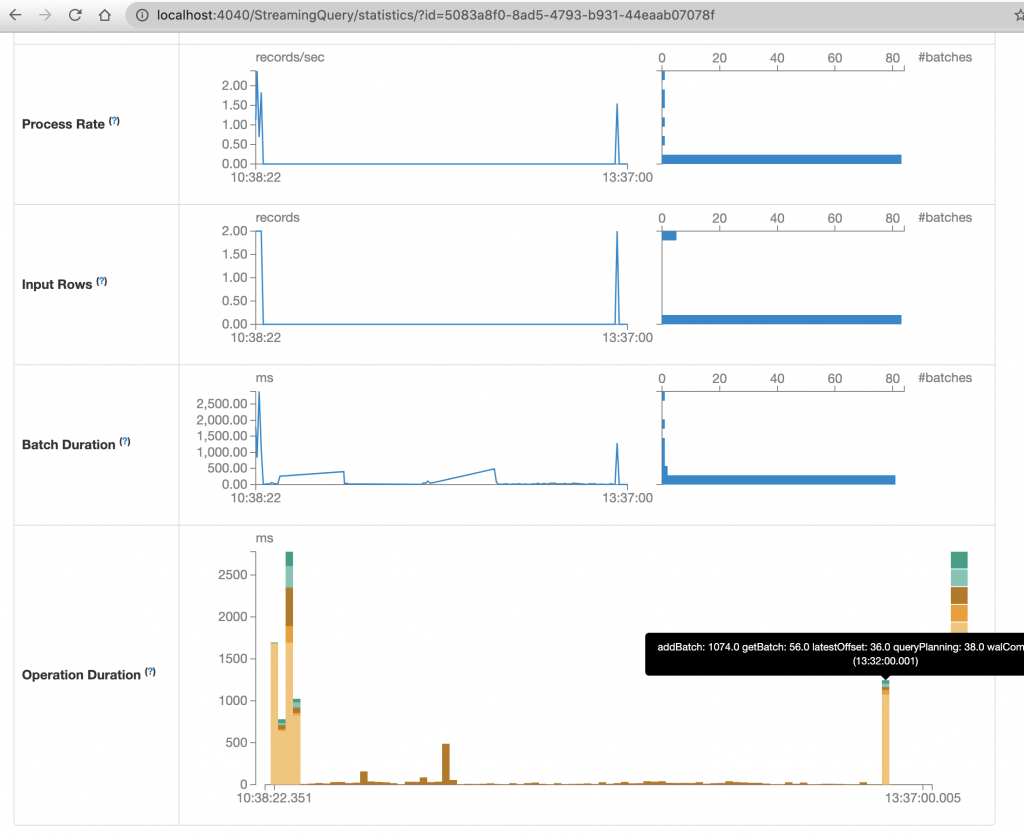
The available information even shows performance related to the rate of processed information which can be a convenient way to improve the performance of the code.
The last chart shows a simple experiment: deleting a folder that stores information of a loaded file makes to reload that file again. The plot shows the operation duration of processing one micro batch. As the code processes only one file per micro batch, this is a simple way to see how much time did the file needed to be ingested.
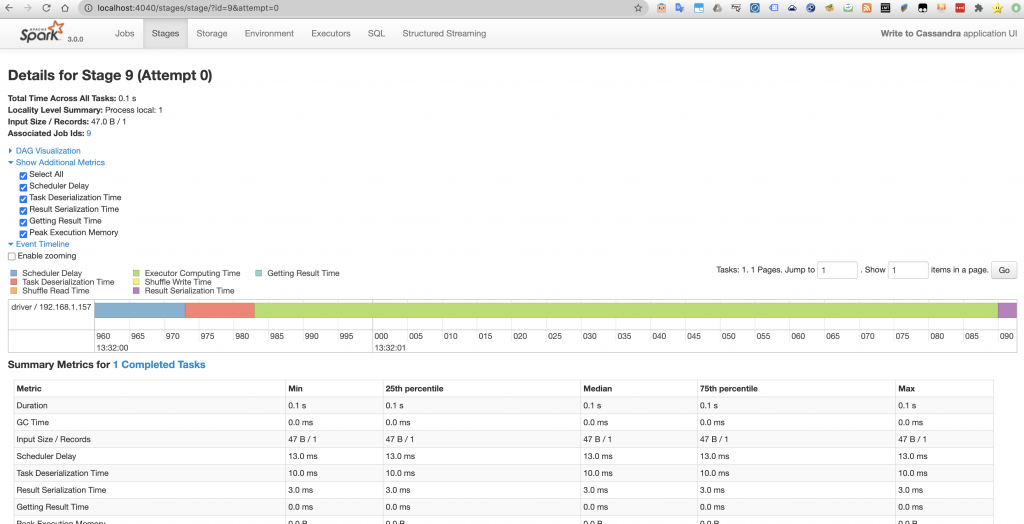
The Spark UI also shows us more detailed information about how the internal operations took place. That is to say what Spark transformations and actions took place, their order and their execution time. This can be a way to see that the Spark calls we made were effectively translated to the transformations and actions that we were expecting.
And this is it for this time!. Shutdown the Cassandra database by stoping the Datastax Desktop stack. Data will remain persistent in the container assigned storage for next time. This will save your computer from remaining with busy resources associated to the Kubernetes cluster that needed to be created. If you are running on top of a laptop watch out the temperatures when running Docker Desktop.
Conclusions
Introducing to Spark Structured Streaming has been relatively easy with the existing possibilities to work on local. I recommend to have a local installation for testing small sets of data. Remember that a speed up in a small dataset translates to big improvement when running in a cluster as data volumes increase so much. I leave open to the reader to try to insert data by using Spark sinks or just employ the same batching technique for other databases that lack a Spark sink. One arguable aspect of spark structured steaming is the way to keep track of the already ingested files. Even if writing data into a disk is a safe way to keep the log of actions it could have been more useful to provide something more practical such as the option to store the ingested files into a fast and lightweight database like RocksDB and exposing that database to the programmer.